Single cell RNA-Sequencing (scRNA-seq) measure transcription activity at single cell resolution, providing novel insights of biological systems and human diseases. Most high-throughput single-cell methods are limited to the quantification of gene-level expression. However, recent advances in long-read sequencing now enable high-throughput sequencing of full-length molecules, opening up an era of single cell isoform-level quantification. New scientific and analytical challenges are presented to properly explore, analyze and document this type of data.
isoswitch is an R package that facilitates the exploration and documentation of isoform expression patterns in long-read scRNA-Seq datasets.
I demonstrate the package functionality in the analysis on two datasets of human nasal epithelium and embryonic lung generated with ScNaUmi-Seq, the long-read scRNA-Seq protocol developed at IPMC (Lebrigand et al, 2020).

|
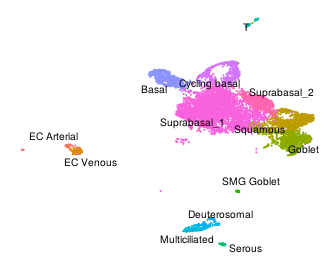 Nasal epithelium |
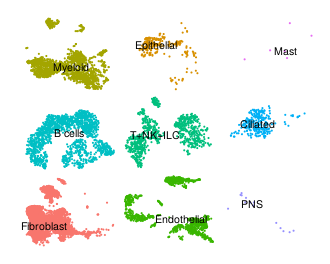 Embryonic lung |

